Our bodies are alive with activity and filled with proteins trapped in fatty membranes or floating in and out of watery cells. Scientists have now, for the first time, captured the dance between the two: a fluid tango involving proteins and fats as they would normally move in cells.
“We’re going beyond taking single snapshots, which give structure but not dynamics, to continuously recording molecules in water, their native state,” says Qian Chen, a materials scientist and engineer at the University of Illinois at Urbana-Champaign. Champaign (UIUC). who led the team and describes their work as ‘filmmaking’.
“We can actually see how proteins change their configuration and, in this case, how the entire self-assembled protein-lipid structure fluctuates over time.”
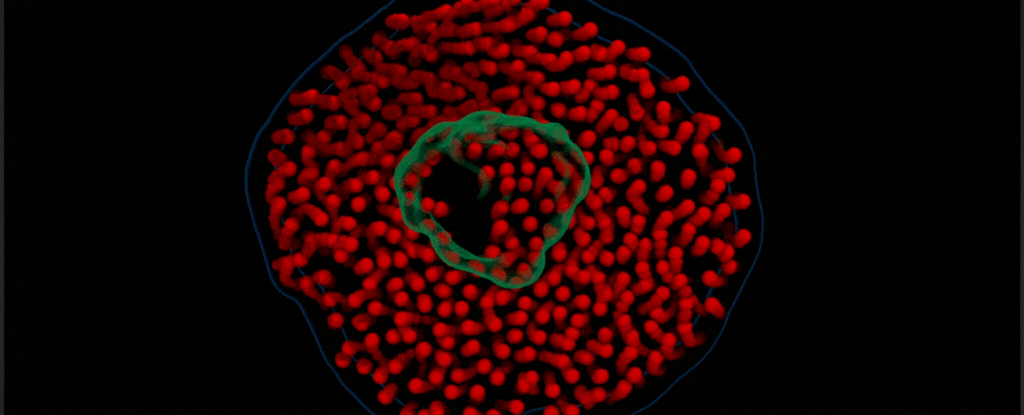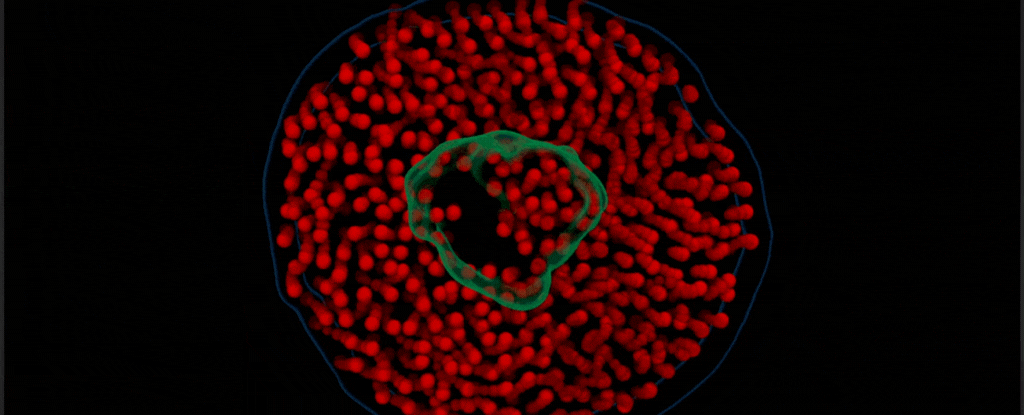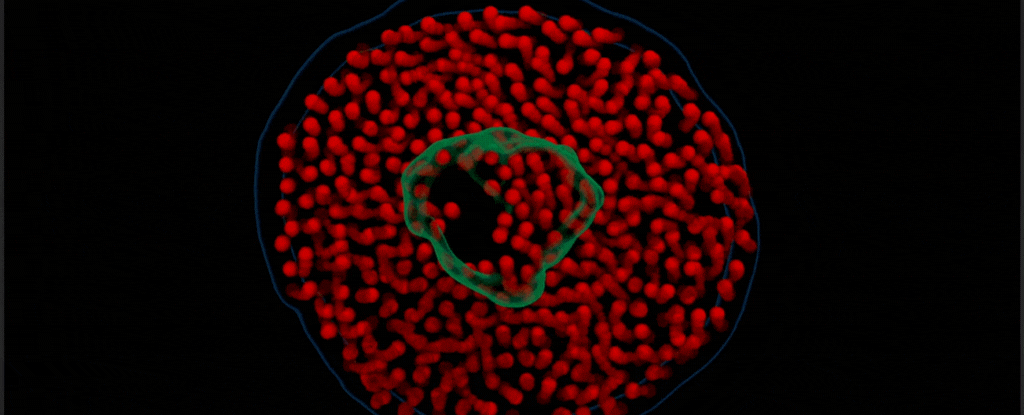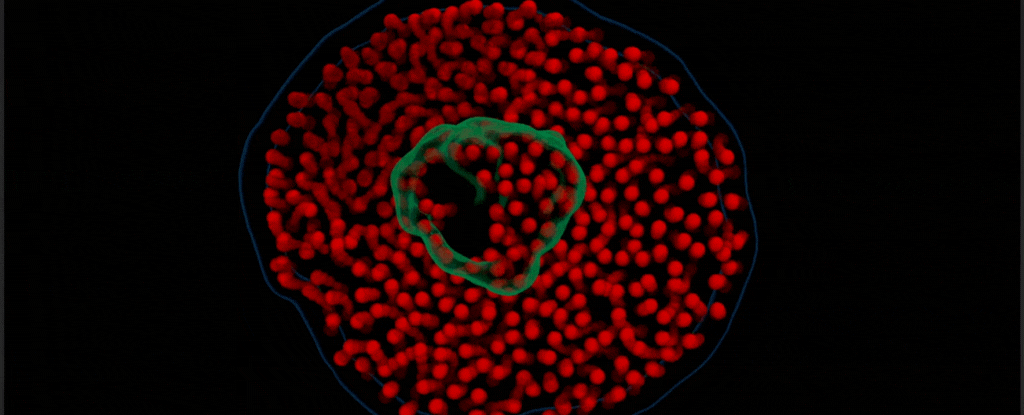
By modifying a widely used imaging technique called transmission electron microscopy, Chen’s team captured the vibrant choreography of membrane protein ‘nanodiscs’ in liquid. These nanodiscs are composed of proteins embedded in a lipid bilayer that resembles the cell membranes in which they are normally found.
The team called their method ‘electronic videography’ and validated the video data by comparing it to atom-level computer models of how molecules should move based on the laws of physics.
The movement of membrane-bound proteins was thought to be quite limited, given how lipids hold them in place. However, the researchers saw interactions between proteins and lipids occurring over much greater distances than previously thought.
Membrane proteins are the cell’s gatekeepers, sensors and signal receivers, so the technique could lead to major advances in our understanding of how they work.
With existing techniques, proteins are usually frozen or crystallized so that they don’t move and blur an image, or be damaged by the X-rays or electron beams used to image them. This gives an inanimate view of a static protein that normally folds and bends, letting scientists understand how it interacts with other molecules based on its structure.
Alternatively, some imaging techniques use a fluorescent molecular tag to track the molecules as they move, rather than looking at the protein directly.
In this case, the researchers placed a drop of water inside two thin sheets of graphene to protect it from the vacuum of the electron microscope. Suspended in the drop of water were nanodiscs of unlabeled proteins and lipids, which the team saw ‘dance’ together as in their natural aqueous environment.
frameborder=”0″ allow=”accelerometer; Play automatically; clipboard-write; encrypted media; gyroscope; picture-in-picture; web-share” referrerpolicy=”strict-origin-when-cross-origin” allowfullscreen>
Materials scientists have tried for at least a decade to film the activity of biological molecules in liquids, but they could not clearly observe the ongoing dynamics of proteins.
With some careful tweaks to the approach, Chen and colleagues imaged their protein-lipid assemblies in real time and in minutes, not microseconds. Importantly, they slowed down the speed of electrons penetrating the sample and worked on the graphene scaffold to successfully film the protein-lipid complex in action.
“Currently, this is really the only experimental way to film this kind of motion over time,” says UIUC materials engineering graduate student John Smith, first author of the paper.
“Life is fluid and it’s in motion. We’re trying to get at the finer details of this connection in an experimental way.”
As for other efforts, improved imaging techniques are revealing incredible detail about all sorts of microscopic events—from watching the outer coat of a virus take shape to capturing the instant proteins collapse into clumps in diseases like Alzheimer’s.
Add artificial intelligence to the mix, to predict the 3D shape of almost every protein known to science, and it sure looks like a new era of biological research has been unlocked.
The research was published in Advances in science.